Model Selection
STA 101L - Summer I 2022
Raphael Morsomme
Welcome
Announcements
- Homework 4 is due on Thursday.
- Homework 5 will be due after the prediction project on Sunday June 5.
- Prediction project
Find your teammate here.
If you have not started yet, start now. You can do already do 80% of the project.
You will have the tools to do the remaining 20% after this lecture.
Recap
simple linear regression model \[ Y \approx \beta_0 + \beta_1 X \]
multiple linear regression model \[ Y \approx \beta_0 + \beta_1 X_1 + \beta_2 X_2 + \dots + + \beta_p X_p \]
categorical predictor
- \((k-1)\) indicator variables
feature engineering
- transforming variables
- combining variables
- interactions
Outline
- Overfitting
- limitations of \(R^2\) for model selection
- Model selection through
- overall criteria (adjusted-\(R^2\), AIC, BIC)
- predictive performance (holdout method, cross-validation)
- stepwise procedure (forward, backward)
Competing models
Competing models
Earlier, we saw that adding the predictor \(\dfrac{1}{\text{displ}}\) gave a better fit.
Let us see if the same idea work with the
treesdataset.
A nonlinear association?
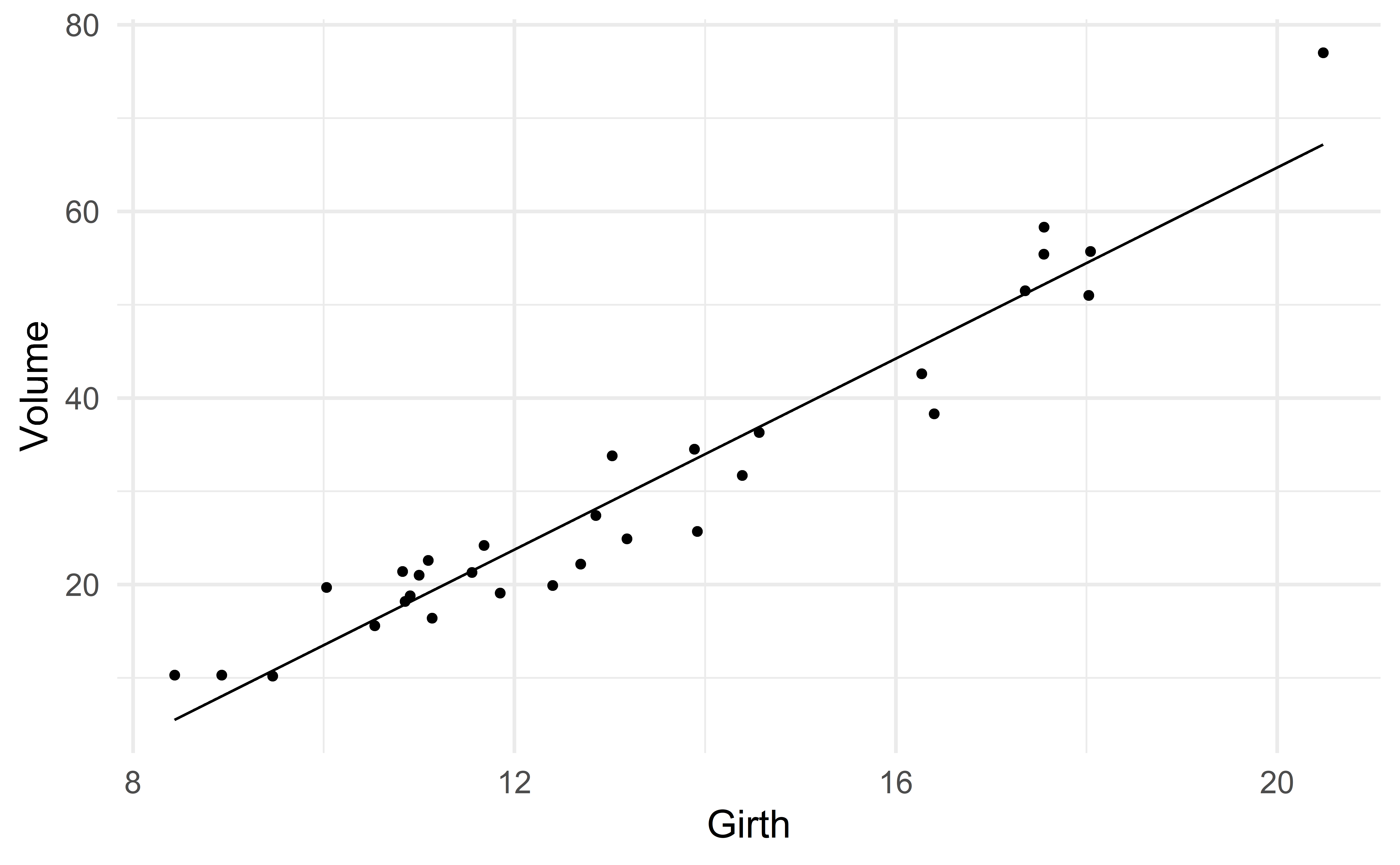
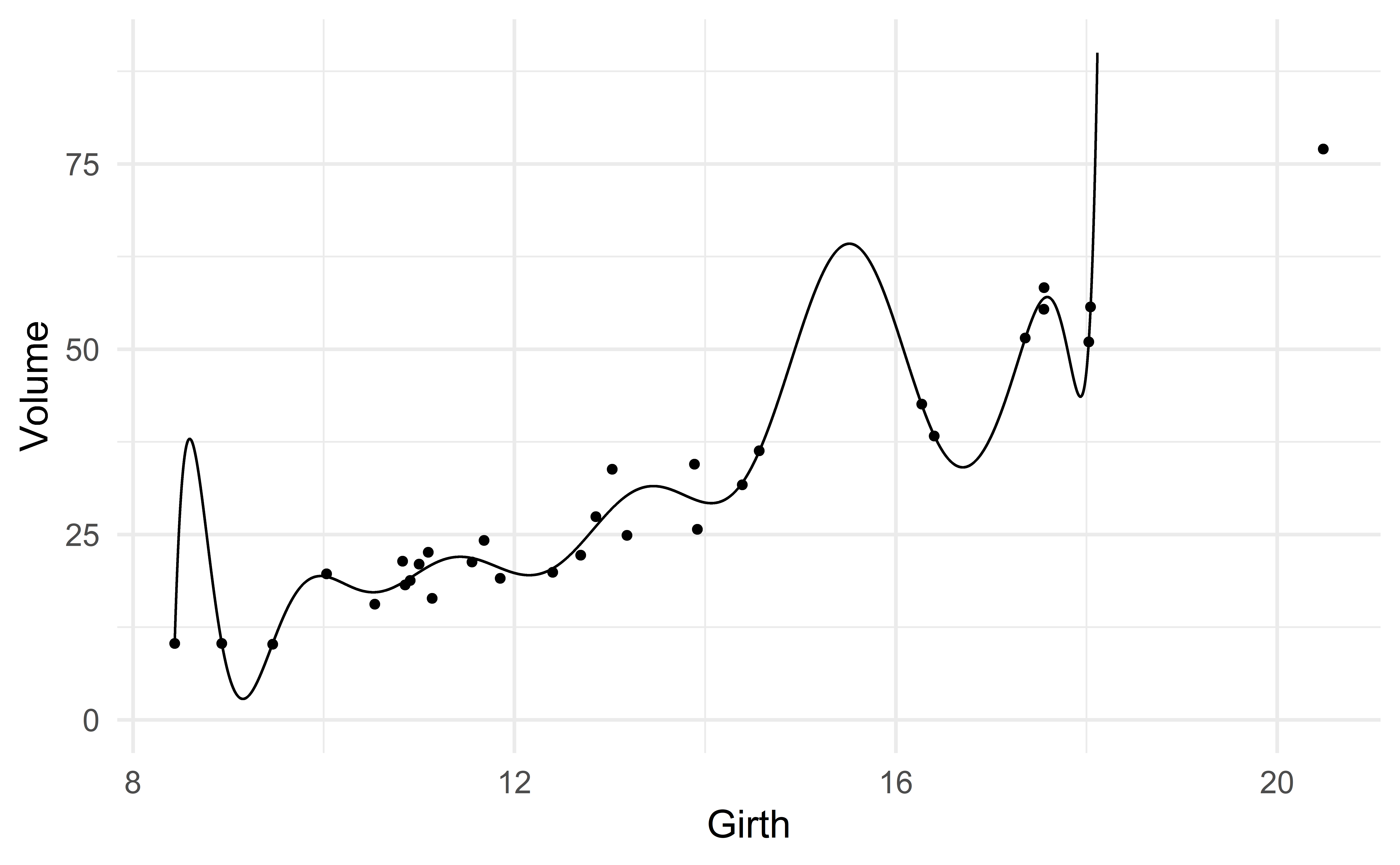
Suppose we want to predict volume using only the variable girth.
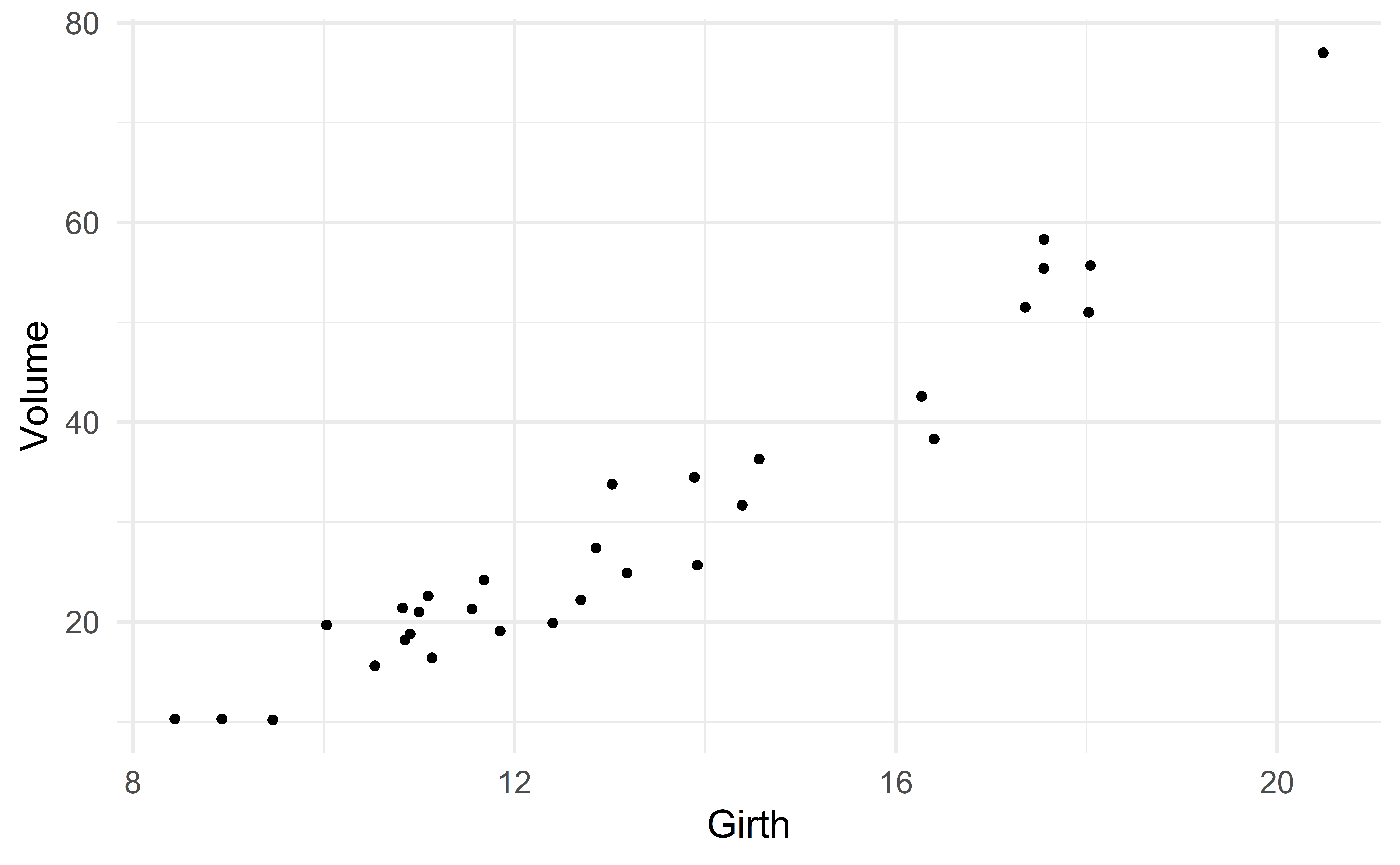
One could argue that there is a slight nonlinear association
Function to compute \(R^2\)
Starting simple…
We start with the simple model
\[ \text{Volume} \approx \beta_0 + \beta_1 \text{girth} \]
Girth_new <- seq(min(d_tree$Girth), max(d_tree$Girth), by = 0.001)
new_data <- tibble(Girth = Girth_new)
Volume_pred <- predict(m1, new_data)
new_data <- mutate(new_data, Volume_pred = Volume_pred)
head(new_data)# A tibble: 6 x 2
Girth Volume_pred
<dbl> <dbl>
1 8.44 5.51
2 8.44 5.52
3 8.44 5.52
4 8.44 5.53
5 8.44 5.53
6 8.44 5.54…taking it up a notch…
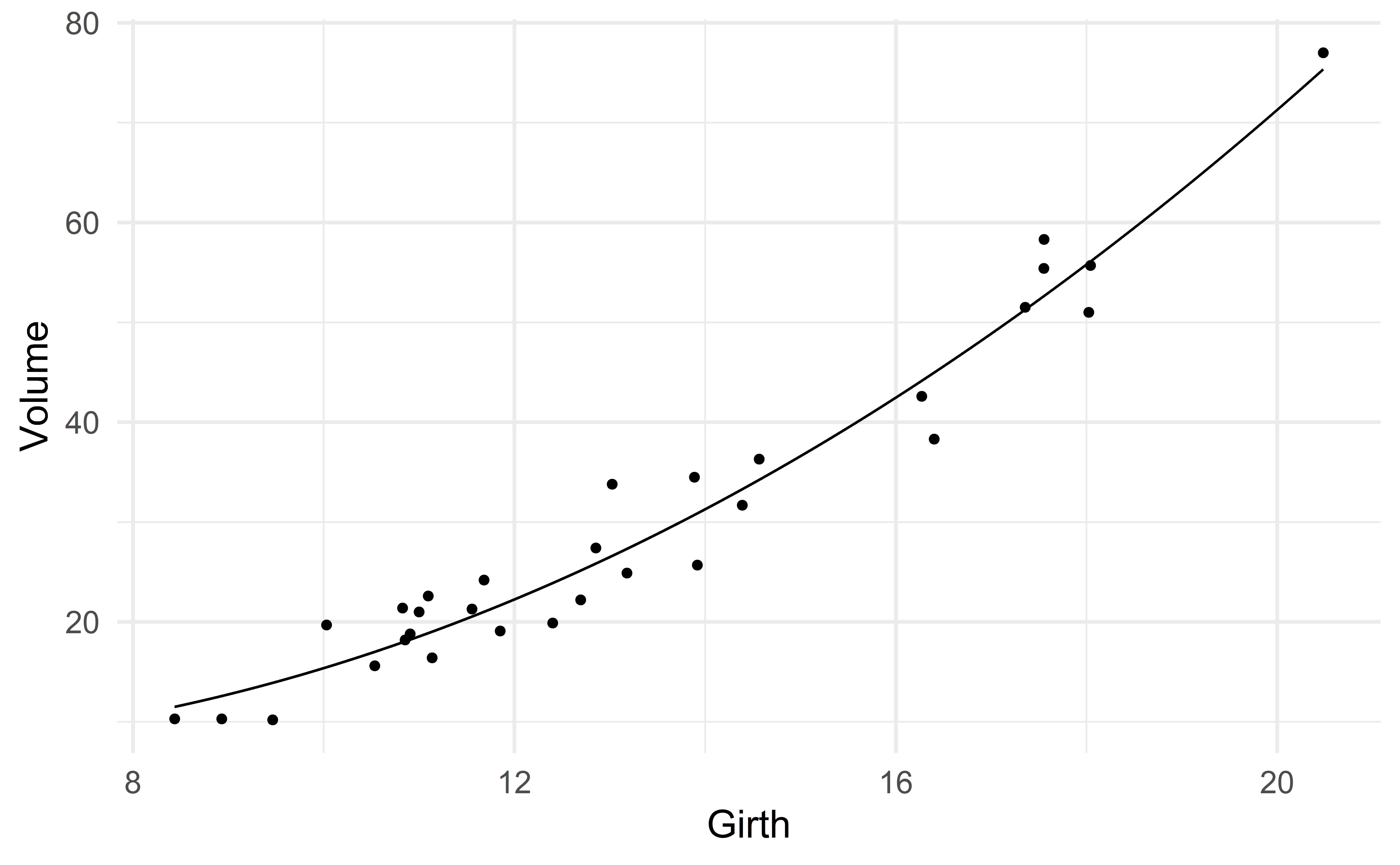
To capture the nonlinear association between Girth and Volume, we consider the predictor \(\text{Girth}^2\).
\[ \text{Volume} \approx \beta_0 + \beta_1 \text{girth} + \beta_2 \text{girth}^2 \]
The command to fit this model is
d_tree2 <- mutate(d_tree, Girth2 = Girth^2)
m2 <- lm(Volume ~ Girth + Girth2, data = d_tree2)
compute_R2(m2)[1] 0.958\(R^2\) has increased! It went from 0.933 (model 1) to 0.958.
Girth_new <- seq(min(d_tree$Girth), max(d_tree$Girth), by = 0.001)
new_data <- tibble(Girth = Girth_new) %>% mutate(Girth2 = Girth^2)
Volume_pred <- predict(m2, new_data)
new_data <- mutate(new_data, Volume_pred = Volume_pred)
head(new_data)# A tibble: 6 x 3
Girth Girth2 Volume_pred
<dbl> <dbl> <dbl>
1 8.44 71.2 11.5
2 8.44 71.2 11.5
3 8.44 71.2 11.5
4 8.44 71.2 11.5
5 8.44 71.2 11.5
6 8.44 71.3 11.5…taking it up another notch…
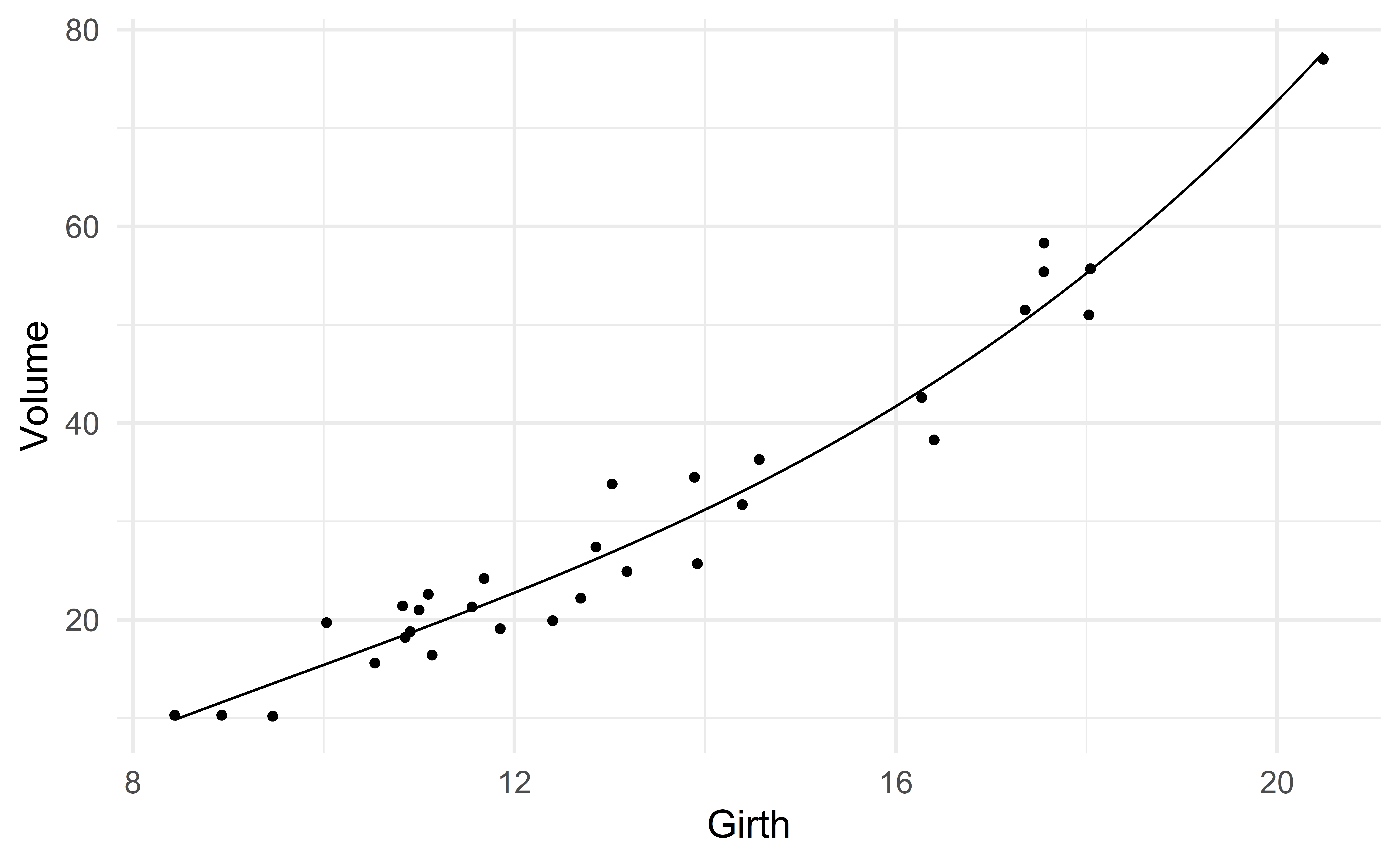
Let us consider the predictor \(\text{Girth}^3\).
\[ \text{Volume} \approx \beta_0 + \beta_1 \text{girth} + \beta_2 \text{girth}^2 + \beta_3 \text{girth}^3 \]
d_tree3 <- mutate(d_tree, Girth2 = Girth^2, Girth3 = Girth^3)
m3 <- lm(Volume ~ Girth + Girth2 + Girth3, data = d_tree3)
compute_R2(m3)[1] 0.96\(R^2\) has increased! It went from 0.958 (model 2) to 0.96.
Girth_new <- seq(min(d_tree$Girth), max(d_tree$Girth), by = 0.001)
new_data <- tibble(Girth = Girth_new) %>% mutate(Girth2 = Girth^2, Girth3 = Girth^3)
Volume_pred <- predict(m3, new_data)
new_data <- mutate(new_data, Volume_pred = Volume_pred)
head(new_data)# A tibble: 6 x 4
Girth Girth2 Girth3 Volume_pred
<dbl> <dbl> <dbl> <dbl>
1 8.44 71.2 601. 9.83
2 8.44 71.2 601. 9.83
3 8.44 71.2 601. 9.83
4 8.44 71.2 601. 9.84
5 8.44 71.2 601. 9.84
6 8.44 71.3 602. 9.84…before taking things to the extreme
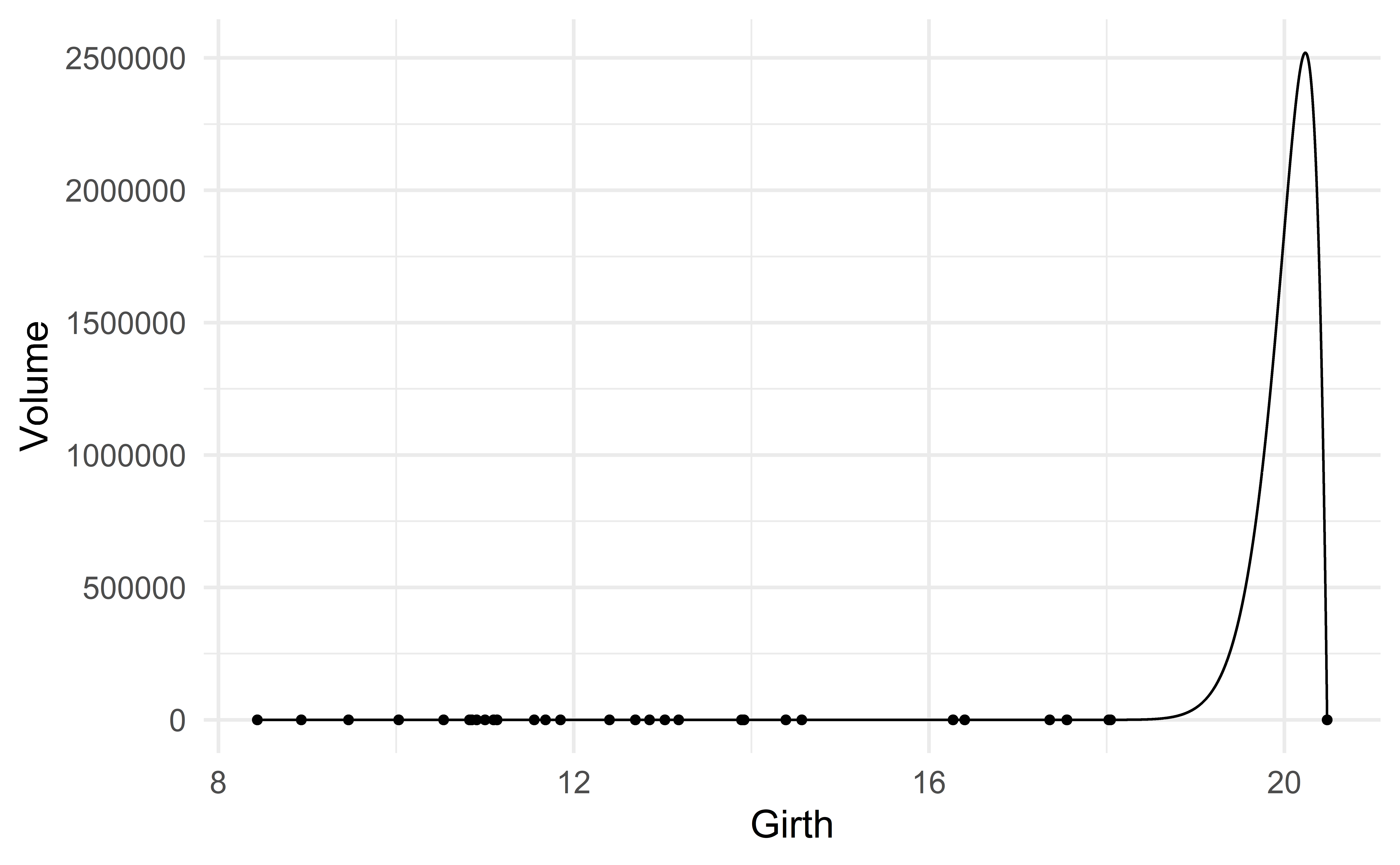
What if we also include the predictors \(\text{Girth}^4, \text{Girth}^5, \dots, \text{Girth}^{k}\) for some larger number \(k\)?
The following R command fits the model with \(k=81\), that is,
\[ \text{Volume} \approx \beta_0 + \beta_1 \text{girth} + \beta_2 \text{girth}^2 + \dots + \beta_{81} \text{girth}^{81} \]
[1] 0.981\(R^2\) has again increased! It went from 0.96 (model 3) to 0.981.
A limitation of \(R^2\)
As we keep adding predictors, \(R^2\) will always increase.
- additional predictors allow the regression line to be more flexible, hence to be closer to the points and reduce the residuals.
Is the model m_extreme a good model?
Note
We want to learn about the relation between Volume and Girth present in the population (not the sample).
A good model should accurately represent the population.
Girth_new <- seq(min(d_tree$Girth), max(d_tree$Girth), by = 0.0001)
new_data <- tibble(Girth = Girth_new)
Volume_pred <- predict(m_extreme, new_data)
new_data <- mutate(new_data, Volume_pred = Volume_pred)
head(new_data)# A tibble: 6 x 2
Girth Volume_pred
<dbl> <dbl>
1 8.44 10.3
2 8.44 10.4
3 8.44 10.4
4 8.44 10.4
5 8.44 10.5
6 8.44 10.5Overfitting
The model m_extreme overfits the data.
A model overfits the data when it fits the sample extremely well but does a poor job for new data.
Remember that we want to learn about the population, not the sample!
Model selection: overall criteria
\(R^2\)
We saw that \(R^2\) keeps increasing as we add predictors.
\[ R^2 = 1 - \dfrac{SSR}{SST} \]
\(R^2\) can therefore not be used to identify models that overfit the data.
Adjusted-\(R^2\)
The adjusted-\(R^2\) is similar to \(R^2\), but penalizes large models:
\[ R^2_{\text{adj}} = 1 - \dfrac{SSR}{SST}\dfrac{n-1}{n-p-1} \]
where \(p\) corresponds to the number of predictors (model size).
The adjusted-\(R^2\) therefore balances goodness of fit and parsimony:
- The ratio \(\dfrac{SSR}{SST}\) favors model with good fit (like \(R^2\))
- The ratio \(\dfrac{n-1}{n-k-1}\) favors parsimonious models
The model with the highest adjusted-\(R^2\) typically provides a good fit without overfitting.
Group exercise - a function for the adjusted-\(R^2\)
Consider the function compute_R2 which takes a model as input and returns its \(R^2\) rounded to the 3rd decimal.
Adapt the function so that it computes the adjusted-\(R^2\) instead of \(R^2\). Give an appropriate name to the new function.
04:00
AIC and BIC
Two other popular overall criteria that balance goodness of fit (small SSR) and parsimony (small \(p\)) are
- the Akaike Information Criterion (AIC)
- the Bayesian Information Criterion (BIC)
The formula for AIC and BIC are respectively
\[ AIC = 2p - \text{GoF}, \qquad BIC = \ln(n)p- \text{GoF} \]
where \(\text{GoF}\) measures the goodness of fit of the model1.
AIC and BIC in practice
Warning
Unlike the adjusted-\(R^2\), smaller is better for the AIC and BIC.
Note
The BIC penalizes the number of parameters \(p\) more strongly than AIC. BIC will therefore tend to favor smaller models (models with fewer parameters).
Computing AIC and BIC
AIC and BIC are easily accessible in R with the command glance.
# A tibble: 4 x 12
r.squared adj.r.squared sigma statistic p.value df logLik AIC BIC
<dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 0.933 0.930 4.34 401. 1.57e-18 1 -88.5 183. 187.
2 0.958 0.955 3.50 317. 5.93e-20 2 -81.3 171. 176.
3 0.960 0.955 3.48 214. 6.39e-19 3 -80.5 171. 178.
4 0.981 0.960 3.28 46.3 2.32e- 9 16 -68.5 173. 199.
# ... with 3 more variables: deviance <dbl>, df.residual <int>, nobs <int>In this case, BIC favor m2, AIC cannot decide between m2 and m3 and the adjusted-\(R^2\) (wrongly) favors m_extreme.
- For AIC and BIC, smaller is better!
- BIC favors more parsimonious models than AIC.
Model selection: predictive performance
Limitations of the previous criteria
The adjusted-\(R^2\), AIC and BIC all try to balance
- goodness of fit
- parsimony
They achieve this balance by favoring models with small SSR while penalizing models with larger \(p\).
…But
- The form of the penalty for \(p\) is somewhat arbitrary, e.g. AIC versus BIC.
- The adjusted-\(R^2\) failed to penalize
m_extreme, although it was clearly overfitting the data.
Predictive performance
Instead of using these criteria, we could look for the model with the best predictions performance.
That is, the model that makes predictions for new observations that are the closest to the true values.
We will learn two approaches to accomplish this
- the holdout method
- cross-validation
The holdout method
The holdout method is a simple method to evaluate the predictive performance of a model.
- Randomly partition your sample into two sets: a training set (typically 2/3 of the sample) and a test set (the remaining 1/3).
- Fit your model to the training set.
- Evaluate the prediction accuracy of the model on the test set.
Note that the test set consists of new observations for the model.
\(\Rightarrow\) Select the model with the best prediction accuracy in step 3.
The holdout method.
Source: IMS
Step 1: training and test sets
The following R function splits a sample into a training and a test set
construct_training_test <- function(sample, prop_training = 2/3){
n <- nrow(sample)
n_training <- round(n * prop_training)
sample_random <- slice_sample(sample, n = n)
sample_training <- slice(sample_random, 1 : n_training )
sample_test <- slice(sample_random, - (1 : n_training))
return(list(
training = sample_training, test = sample_test
))
} Girth Height Volume
1 8.931477 70 10.3
2 8.436883 65 10.3
3 9.464900 63 10.2
4 11.136215 72 16.4
5 10.907321 81 18.8
6 10.030025 83 19.7
7 10.535716 66 15.6
8 10.852640 75 18.2
9 11.097116 80 22.6
10 12.402327 75 19.9
11 11.681797 79 24.2
12 11.000495 76 21.0
13 10.826171 76 21.4
14 11.555269 69 21.3
15 11.850392 75 19.1
16 12.694245 74 22.2
17 13.026112 85 33.8
18 12.854039 86 27.4
19 13.917842 71 25.7
20 13.181231 64 24.9
21 13.887866 78 34.5
22 14.388698 80 31.7
23 14.566668 74 36.3
24 16.402095 72 38.3
25 16.271447 77 42.6
26 17.551804 81 55.4
27 18.042885 82 55.7
28 17.554523 80 58.3
29 17.357700 80 51.5
30 18.023363 80 51.0
31 20.482147 87 77.0set.seed(0) # set the seed of the random number generator to 0
training_test_sets <- construct_training_test(d_tree)
training_set <- training_test_sets[["training"]]
training_set Girth Height Volume
1 11.555269 69 21.3
2 16.271447 77 42.6
3 11.136215 72 16.4
4 10.535716 66 15.6
5 8.931477 70 10.3
6 8.436883 65 10.3
7 14.566668 74 36.3
8 11.681797 79 24.2
9 20.482147 87 77.0
10 12.854039 86 27.4
11 13.917842 71 25.7
12 18.042885 82 55.7
13 12.402327 75 19.9
14 18.023363 80 51.0
15 13.887866 78 34.5
16 17.554523 80 58.3
17 11.097116 80 22.6
18 10.907321 81 18.8
19 14.388698 80 31.7
20 11.850392 75 19.1
21 11.000495 76 21.0Step 2: fit the model to the training set
We simply fit our regression model to the training set.
Step 3: Evaluate the prediction accuracy on the test set
To evaluate the prediction accuracy, we start by computing the predictions for the observations in the test set.
A good model will make predictions that are closed to the true values of the response variable.
Sum of squared errors
A good measure of prediction accuracy is the sum of squared errors (SSE).
\[ SSE = (y_1 - \hat{y}_1)^2 + (y_2 - \hat{y}_2)^2 + \dots + (y_m - \hat{y}_m)^2 \]
Small SSE is better.
Mean squared error
In practice, the (root) mean sum of squared errors ((R)MSE) is often used.
\[ MSE = \dfrac{SSE}{m} = \dfrac{(y_1 - \hat{y}_1)^2 + (y_2 - \hat{y}_2)^2 + \dots + (y_m - \hat{y}_m)^2}{m} \]
\[ RMSE = \sqrt{\dfrac{SSE}{m}} = \sqrt{\dfrac{(y_1 - \hat{y}_1)^2 + (y_2 - \hat{y}_2)^2 + \dots + (y_m - \hat{y}_m)^2}{m}} \] Again, small (R)MSE is better.
SSE <- sum ((Volume - Volume_hat)^2)
MSE <- mean((Volume - Volume_hat)^2)
RMSE <- sqrt(MSE)
SSE; MSE; RMSE[1] 211.4543[1] 21.14543[1] 4.598416(R)MSE
The advantage of (R)MSE over SSE is that we can compare models that are evaluated with test sets of different sizes.
Selecting a model
Apply steps 2 and 3 on different models.
- use the same training and test sets from step 1 for the different models
\(\Rightarrow\) Choose the model with the lowest (R)MSE!
This model has the best predictive accuracy.
Function to compute the RMSE
# Step 1 - partition sample into a training set and a test set
# already done
# Step 2 - fit models to training set
m1 <- lm(Volume ~ poly(Girth, degree = 1 , raw = TRUE), data = training_set)
m2 <- lm(Volume ~ poly(Girth, degree = 2 , raw = TRUE), data = training_set)
m3 <- lm(Volume ~ poly(Girth, degree = 3 , raw = TRUE), data = training_set)
m48 <- lm(Volume ~ poly(Girth, degree = 48, raw = TRUE), data = training_set)
# Step 3 - evaluate models on test set
compute_RMSE(test_set, m1)
compute_RMSE(test_set, m2)
compute_RMSE(test_set, m3) # best
compute_RMSE(test_set, m48)[1] 4.598416
[1] 4.07333
[1] 3.957379
[1] 29.97533We select m3.
Limitation of the holdout method
The previous analysis was conducted with set.seed(0).
Note that models m2 and m3 were pretty close.
Could we have obtained a different result with a different seed?
# Step 1
set.seed(1) # new seed
training_test_sets <- construct_training_test(d_tree)
training_set <- training_test_sets[["training"]]
test_set <- training_test_sets[["test"]]
# Step 2
m1 <- lm(Volume ~ poly(Girth, degree = 1 , raw = TRUE), data = training_set)
m2 <- lm(Volume ~ poly(Girth, degree = 2 , raw = TRUE), data = training_set)
m3 <- lm(Volume ~ poly(Girth, degree = 3 , raw = TRUE), data = training_set)
m48 <- lm(Volume ~ poly(Girth, degree = 48, raw = TRUE), data = training_set)
# Step 3
compute_RMSE(test_set, m1)
compute_RMSE(test_set, m2) # best
compute_RMSE(test_set, m3)
compute_RMSE(test_set, m48)[1] 6.097541
[1] 4.248172
[1] 10.37698
[1] 103753769The test set matters!
With set.seed(1), the holdout method indicates that m2 is the best model.
m3, our previous best model, is worse thanm1!
A drawback of the holdout method is that the test set matters a lot.
- Repeating steps 2 and 3 with a different partition in step 1 may give different results.
Group exercise - limitation of the holdout method
Copy and paste the following code in your R session. Try a few different seed numbers for step 1. Keep track of which model has the lowest RMSE. Is m1 ever the best model?
You can safely ignore the warning messages.
# Setup
set.seed(0) # do not change this seed number
d_tree <- datasets::trees %>% mutate(Girth = Girth + rnorm(nrow(.), 0, 0.5))
compute_RMSE <- function(test_set, model){
y <- test_set$Volume
y_hat <- predict(model, test_set)
E <- y - y_hat
MSE <- mean(E^2)
RMSE <- sqrt(MSE)
return(RMSE)
}
construct_training_test <- function(sample, prop_training = 2/3){
n <- nrow(sample)
n_training <- round(n * prop_training)
sample_random <- slice_sample(sample, n = n)
sample_training <- slice(sample_random, 1 : n_training )
sample_test <- slice(sample_random, - (1 : n_training))
return(list(
training = sample_training, test = sample_test
))
}
# Step 1
set.seed(2) # try a few different seed numbers
training_test_sets <- construct_training_test(d_tree)
training_set <- training_test_sets[["training"]]
test_set <- training_test_sets[["test"]]
# Step 2
m1 <- lm(Volume ~ poly(Girth, degree = 1 , raw = TRUE), data = training_set)
m2 <- lm(Volume ~ poly(Girth, degree = 2 , raw = TRUE), data = training_set)
m3 <- lm(Volume ~ poly(Girth, degree = 3 , raw = TRUE), data = training_set)
m48 <- lm(Volume ~ poly(Girth, degree = 48, raw = TRUE), data = training_set)
# Step 3
compute_RMSE(test_set, m1)
compute_RMSE(test_set, m2)
compute_RMSE(test_set, m3)
compute_RMSE(test_set, m48)05:00
Cross-validation
Cross-validation (CV) is a natural generalization of the holdout method:
- repeat the holdout method many times with different partitions in step 1.
- randomly partition the sample into \(k\) folds of equal size. \(k\) is typically \(5\) or \(10\).
Repeat the following steps \(k\) times
Let fold 1 be the test set and the remaining folds be the training set.
Fit the model to the training set (like the holdout method)
Evaluate the prediction accuracy of the model on the test set (like the holdout method)
Go back to step 1, let the next fold be the test set.
\(\Rightarrow\) Select the model with the best overall prediction accuracy in step 3.
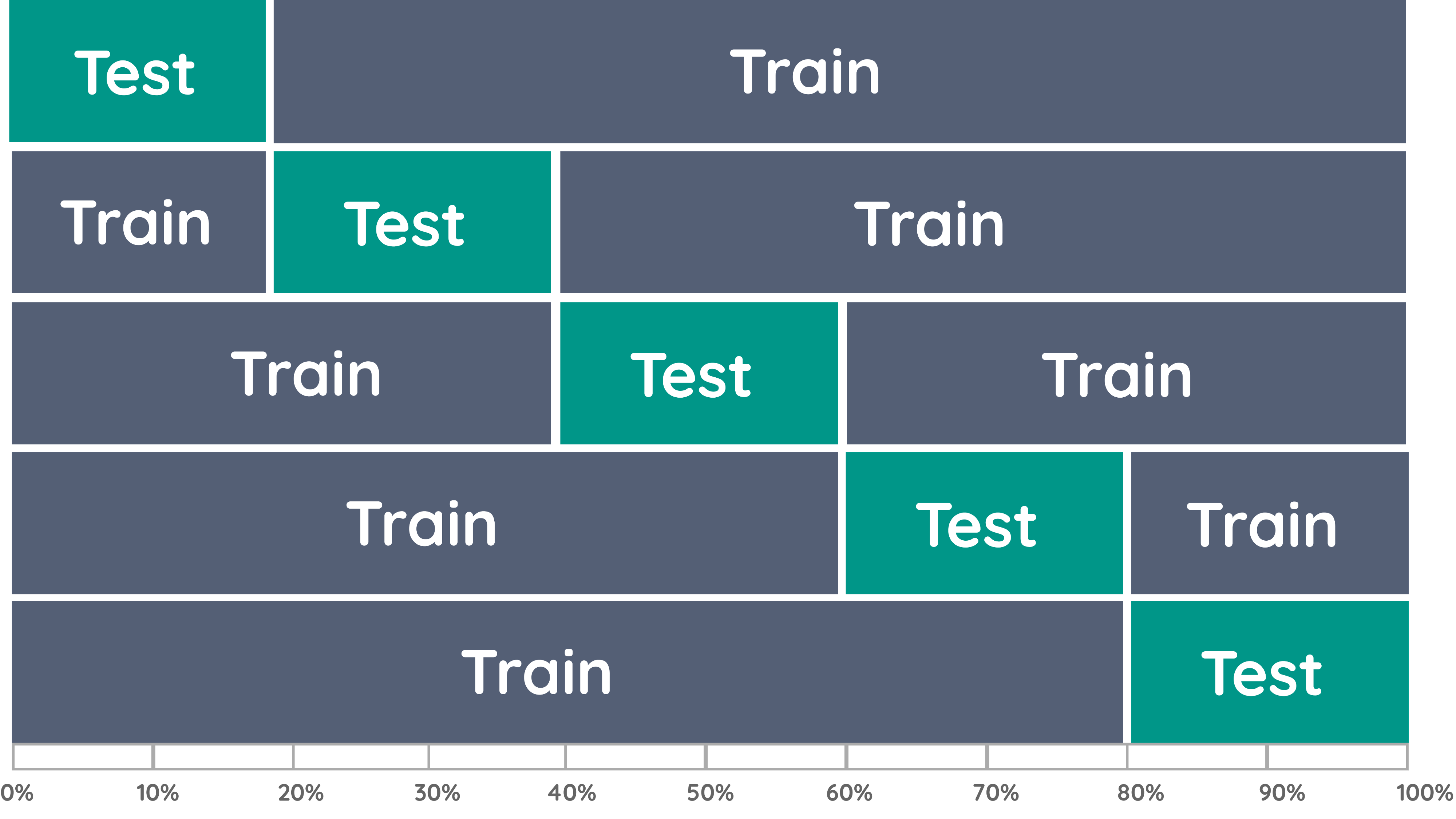
Source: towardsdatascience
Step 0
set.seed(0)
# Setup
n <- nrow(d_tree)
n_folds <- 10
# Fold assignment
folds <- c(rep(1:n_folds, n %/% n_folds), seq_along(n %% n_folds))
# you do not need to understand the details of the previous line
# just note that it creates folds of (almost) the same size
folds [1] 1 2 3 4 5 6 7 8 9 10 1 2 3 4 5 6 7 8 9 10 1 2 3 4 5
[26] 6 7 8 9 10 1# Step 1
d_tree_fold <- d_tree %>%
slice_sample(n = n) %>% # shuffle the rows
mutate(fold = folds)
head(d_tree_fold) Girth Height Volume fold
1 11.555269 69 21.3 1
2 16.271447 77 42.6 2
3 11.136215 72 16.4 3
4 10.535716 66 15.6 4
5 8.931477 70 10.3 5
6 8.436883 65 10.3 6Steps 1-4 – Setup
Steps 1-4
RMSE <- create_empty_RMSE()
for(i in 1 : n_folds){
# Step 1
training_set <- filter(d_tree_fold, fold != i)
test_set <- filter(d_tree_fold, fold == i)
# Step 2
m1 <- lm(Volume ~ poly(Girth, degree = 1 , raw = TRUE), data = training_set)
m2 <- lm(Volume ~ poly(Girth, degree = 2 , raw = TRUE), data = training_set)
m3 <- lm(Volume ~ poly(Girth, degree = 3 , raw = TRUE), data = training_set)
m48 <- lm(Volume ~ poly(Girth, degree = 48, raw = TRUE), data = training_set)
# Step 3
rmse_m1 <- compute_RMSE(test_set, m1 )
rmse_m2 <- compute_RMSE(test_set, m2 )
rmse_m3 <- compute_RMSE(test_set, m3 )
rmse_m48 <- compute_RMSE(test_set, m48)
RMSE <- RMSE %>%
add_row(fold = i, rmse_m1, rmse_m2, rmse_m3, rmse_m48)
}Model selection with RMSE
Which model has the lowest RMSE across all folds?
# A tibble: 10 x 5
fold rmse_m1 rmse_m2 rmse_m3 rmse_m48
<dbl> <dbl> <dbl> <dbl> <dbl>
1 1 5.07 3.50 3.54 4.25
2 2 2.89 2.34 1.95 10.8
3 3 4.78 5.02 5.11 5.54
4 4 2.82 3.49 3.45 3.15
5 5 1.67 2.99 2.67 72.5
6 6 6.38 4.45 5.38 412.
7 7 3.52 2.89 2.83 4.73
8 8 1.37 2.01 1.77 3.90
9 9 7.49 3.14 3.92 2481852.
10 10 5.60 4.53 4.27 4.47# A tibble: 1 x 5
fold rmse_m1 rmse_m2 rmse_m3 rmse_m48
<dbl> <dbl> <dbl> <dbl> <dbl>
1 5.5 4.16 3.44 3.49 248237.# A tibble: 1 x 5
fold rmse_m1 rmse_m2 rmse_m3 rmse_m48
<dbl> <dbl> <dbl> <dbl> <dbl>
1 55 41.6 34.4 34.9 2482374.m2 has slightly better overall RMSE.
04:00
Short podcast on cross-validation
Cross-validation is a central technique in machine learning.
Still not totally clear how it works? Check out this podcast to hear more about this important topic.
These two other podcasts on muliple linear regression and \(R^2\) are also excellent.
Taking things to the extreme: LOOCV
Set \(k=n\).
that is, use \(n\) folds, each of size \(1\).
Each test set will therefore consist of a single observation and the corresponding training set of the remaining \(n-1\) observations.
LOOCV
RMSE <- create_empty_RMSE()
# Step 0
n <- nrow(d_tree)
d_tree_loocv <- mutate(d_tree, folds = 1 : n) # every observation belongs to a different fold
for(i in 1:n){
# Step 1
training_set <- filter(d_tree_loocv, folds != i)
test_set <- filter(d_tree_loocv, folds == i)
# Step 2
m1 <- lm(Volume ~ poly(Girth, degree = 1 , raw = TRUE), data = training_set)
m2 <- lm(Volume ~ poly(Girth, degree = 2 , raw = TRUE), data = training_set)
m3 <- lm(Volume ~ poly(Girth, degree = 3 , raw = TRUE), data = training_set)
m48 <- lm(Volume ~ poly(Girth, degree = 48, raw = TRUE), data = training_set)
# Step 3
rmse_m1 <- compute_RMSE(test_set, m1 )
rmse_m2 <- compute_RMSE(test_set, m2 )
rmse_m3 <- compute_RMSE(test_set, m3 )
rmse_m48 <- compute_RMSE(test_set, m48)
RMSE <- RMSE %>%
add_row(fold = i, rmse_m1, rmse_m2, rmse_m3, rmse_m48)
}
summarize_all(RMSE, mean)# A tibble: 1 x 5
fold rmse_m1 rmse_m2 rmse_m3 rmse_m48
<dbl> <dbl> <dbl> <dbl> <dbl>
1 16 3.64 3.11 3.12 142697.Model selection: stepwise procedures
Stepwise procedures
Not my favorite method, but it is widely used.
- Akin the “throw cooked spaghetti to the wall and see what sticks” technique.
- This is what you do when you are clueless about what variables may be good predictors
- …but then you should probably collaborate with a scientific expert.
Two types of stepwise procedures: (i) forward selection and (ii) backward elimination.
Forward selection
Choose an overall criterion that balances GoF (small SSR) and parsimony (small \(p\))
\(\Rightarrow\) adjusted-\(R^2\), AIC or BIC
Start with the empty model \(Y \approx \beta_0\), i.e. the model with no predictor. This is our current model
Fit all possible models with one additional predictor.
Compute the criterion, say the AIC, of each of these models.
Identify the model with the smallest AIC. This is our candidate model.
If the AIC of the candidate model is smaller (better) than the AIC of the current model, the candidate model becomes the current model. Go back to step 3.
If the AIC of the candidate model is larger than the AIC of the current model (no new model improves on the current one), the procedure stops, and we select the current model.
Group exercise - forward selection
Exercise 8.13. In addition, fit the candidate model (a single model) in R with the command lm.
05:00
Backward elimination
Similar to forward selection, except that
- We start with the full model.
- We remove one predictor at a time,
- until removing any predictor makes the model worse.
Group exercise - backward elimination
Exercise 8.11. In addition, fit the current and candidate models in R with the command lm.
05:00
Forward selection versus backward elimination
Note that forward selection and backward elimination need not agree; they may select different models!
. . .
- What to do in that case? Nobody knows.
How to prevent overfitting?
Parsimony
To obtain a parsimonious model
- use your knowledge of the subject to carefuly choose which variables to consider and construct new ones, and
- implement a model selection procedure.
Recap
Recap
Overfitting
- limitations of \(R^2\) for model selection
Model selection through
overall criteria (adjusted-\(R^2\), AIC, BIC)
predictive performance (holdout method, cross-validation)
stepwise procedure (forward, backward)
You are now fully equipped for the prediction project!
🍀 Good luck! 🍀